Some recent publications
Juul*, J. S., Jensen, M. H. & Krishna*, S. (2019) Constraints on somite formaiton in developing embryos. J. R. Soc. Interface 16, doi:10.1098/rsif.2019.0451 (*Co-corresponding authors)Varahan, S., Sinha, V., Walwekar, A., Krishna, S. & Laxman, S. (2019) Metabolic constraints drive self-organization of specialized cell groups, eLife 8, e46735
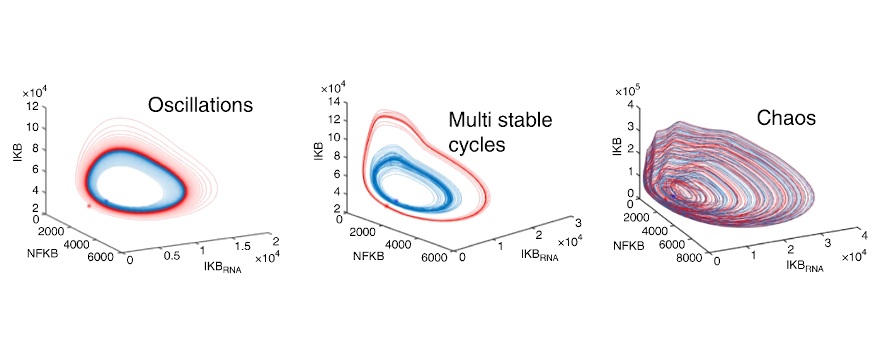
Heltberg, M., Krishna*, S. & Jensen*, M. H. (2019) Chaotic dynamics of transcription factors and its consequences for downstream gene expression, Nat. Commun. 10, 71 (*Co-corresponding authors)
Krishna, S & Laxman, S. (2018) A minimal "push-pull" bistability model explains oscillations between quiescent and proliferative cell states, Mol. Biol. Cell 29, 2243-2258
Suratekar, R., Panda, A., Raghu, P. & Krishna, S. (2018) Evidence of sinks and sources in the plc activated pip2 cycle, FEBS Lett. 592, 962–972
Datta, V., Siddharthan, R. & Krishna, S. (2018) Detection of cooperatively bound transcription factor pairs using chip-seq peak intensities and expectation maximization, PLoS ONE 13, e0199771
Lal, A., Krishna, S. & Seshasayee, A. S. N. (2018) Regulation of global transcription in e. coli by rsd and 6s rna, G3 8, 2079–2089
Sinha, V., Goyal, A., Svenningsen, S. L., Semsey, S. & Krishna, S. (2017) In silico evolution of lysis-lysogeny strategies reproduces observed lysogeny propensities in temperate bacteriophages, Front. Microbiol. 8, 1386.
Heltberg, M., Kellogg, R. A., Krishna, S., Tay, S. & Jensen, M. H. (2016) Noise induces hopping between nf-kb entrainment modes, Cell Sys. 3, 532.
Full publication list
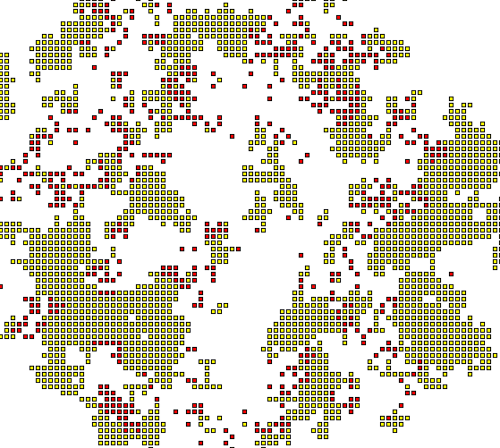
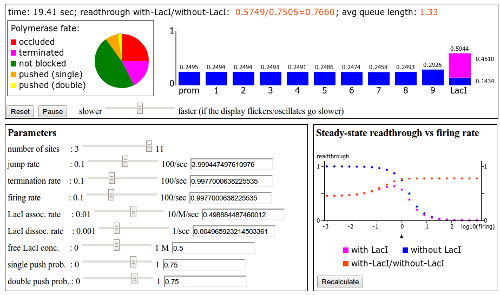
Interactive models
Talks & videos
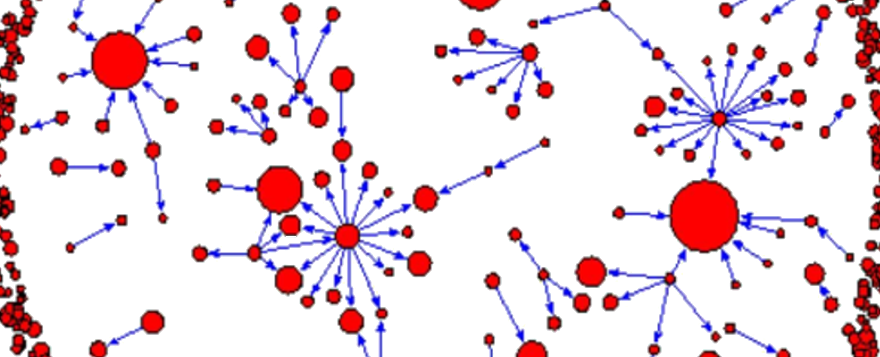
Today Dr. Sandeep Krishna introduces the subject of Complex Networks to the #MonsoonSchool. A bit about what that means: pic.twitter.com/hc1RUCCeod
— National Centre for Biological Sciences (@NCBS_Bangalore) June 16, 2018